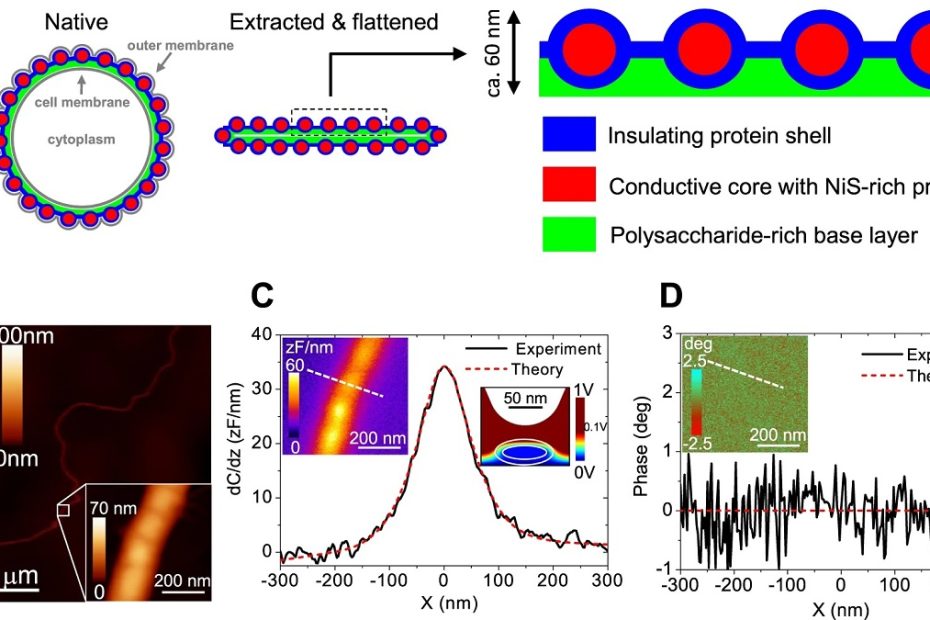
Efficient long-range conduction in cable bacteria through nickel protein wires
Bio-materials typically have an intrinsically low electrical conductivity, and so the availability of a bio-material with extraordinary electrical properties has great potential for new applications… Read More »Efficient long-range conduction in cable bacteria through nickel protein wires